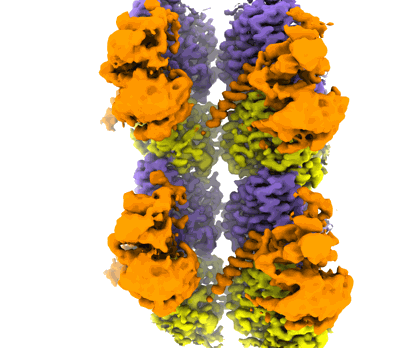
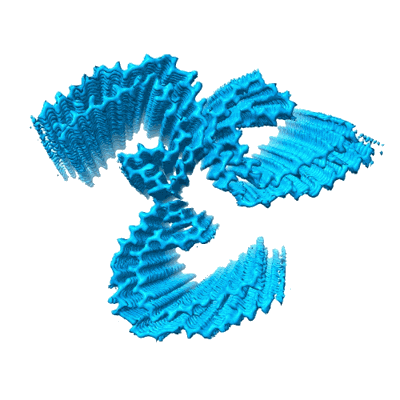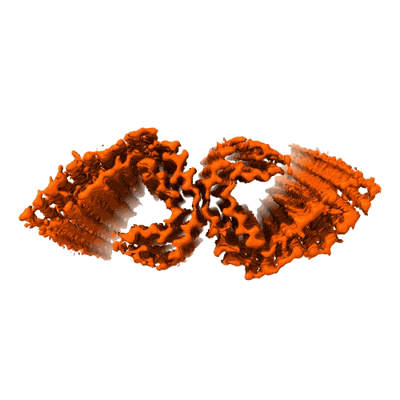Mahalingan KK, Grotjahn DA, Li Y, Lander GC, Zehr EA, Roll-Mecak A
Nat Chem Biol (2024) [ DOI: doi:10.1038/s41589-024-01599-0 Pubmed: 38658656 ]
- Tubulin beta chain (49 kDa, Protein from Homo sapiens)
- Tubulin polyglutamylase ttll6 (Complex from Mus musculus)
- Magnesium ion (24 Da, Ligand)
- Ttll6 bound to unmodified human microtubules (300 kDa, Complex)
- Phosphomethylphosphonic acid guanylate ester (521 Da, Ligand)
- Tubulin alpha-1b chain, tubulin betai chain (Complex from Homo sapiens)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Tubulin alpha-1b chain (50 kDa, Protein from Homo sapiens)
- Tubulin polyglutamylase ttll6 (59 kDa, Protein from Mus musculus)
J Biol Chem (2024) pp. 107326-107326 [ DOI: doi:10.1016/j.jbc.2024.107326 Pubmed: 38679331 ]
PHF filament generated from 4E-Tau(297-407) under neutral Mg2+ condition
J Biol Chem (2024) pp. 107326-107326 [ DOI: doi:10.1016/j.jbc.2024.107326 Pubmed: 38679331 ]
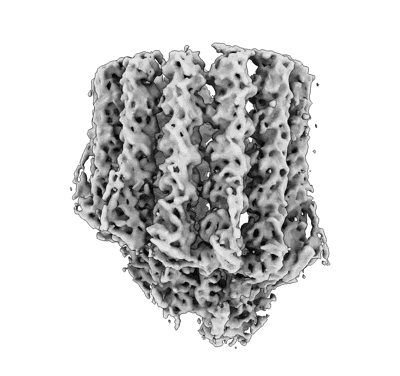
TUBB4B and TUBA1A Heterodimer from Human Respiratory Doublet Microtubules
To Be Published
- Tubulin beta-4b chain (49 kDa, Protein from Homo sapiens)
- Magnesium ion (24 Da, Ligand)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Tubb4b and tuba1a heterodimer from human respiratory doublet microtubules (99 kDa, Complex from Homo sapiens)
- Guanosine-5'-diphosphate (443 Da, Ligand)
- Tubulin alpha-1a chain (50 kDa, Protein from Homo sapiens)
Resolution: 4.3 Å
EM Method: Single-particle
Fitted PDBs: 8v3l
Q-score: 0.151
To Be Published
- Ccdc81b (179 kDa, Protein from Tetrahymena thermophila)
- 48-nm proximal repeat unit of microtubule doublet from tetrahymena thermophila strain k40r (7 MDa, Cellular component from Tetrahymena thermophila)
- Bmip1 (28 kDa, Protein from Tetrahymena thermophila)
Structure of the native microtubule lattice nucleated from the yeast spindle pole body
Resolution: 6.6 Å
EM Method: Subtomogram averaging
Fitted PDBs: 8qv0
Q-score: 0.123
Dendooven T, Yatskevich S, Burt A, Chen ZA, Bellini D, Rappsilber J, Kilmartin JV, Barford D
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01281-y Pubmed: 38609662 ]
Resolution: 9.2 Å
EM Method: Subtomogram averaging
Fitted PDBs: 8qv2
Q-score: 0.112
Dendooven T, Yatskevich S, Burt A, Chen ZA, Bellini D, Rappsilber J, Kilmartin JV, Barford D
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01281-y Pubmed: 38609662 ]
- Water (18 Da, Ligand)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Y-tubulin ring complex capping the microtubule minus end (Cellular component from Saccharomyces cerevisiae)
- Tubulin gamma chain (52 kDa, Protein from Saccharomyces cerevisiae)
- Spindle pole body component (98 kDa, Protein from Saccharomyces cerevisiae)
- Tubulin alpha-1 chain (49 kDa, Protein from Saccharomyces cerevisiae)
- Spindle pole body component 110 (111 kDa, Protein from Saccharomyces cerevisiae)
- Guanosine-5'-diphosphate (443 Da, Ligand)
- Spindle pole body component (96 kDa, Protein from Saccharomyces cerevisiae)
- Unkown protein (5 kDa, Protein from Saccharomyces cerevisiae)
- Tubulin beta chain (50 kDa, Protein from Saccharomyces cerevisiae)
Resolution: 8.2 Å
EM Method: Subtomogram averaging
Fitted PDBs: 8qv3
Q-score: 0.144
Dendooven T, Yatskevich S, Burt A, Chen ZA, Bellini D, Rappsilber J, Kilmartin JV, Barford D
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01281-y Pubmed: 38609662 ]
- Water (18 Da, Ligand)
- Unknown protein (5 kDa, Protein from Saccharomyces cerevisiae)
- Guanosine-5'-triphosphate (523 Da, Ligand)
- Tubulin gamma chain (52 kDa, Protein from Saccharomyces cerevisiae)
- Spindle pole body component 110 (111 kDa, Protein from Saccharomyces cerevisiae)
- Spindle pole body component (98 kDa, Protein from Saccharomyces cerevisiae)
- Tubulin alpha-1 chain (49 kDa, Protein from Saccharomyces cerevisiae)
- Tubulin beta chain (50 kDa, Protein from Saccharomyces cerevisiae)
- Spindle pole body component (96 kDa, Protein from Saccharomyces cerevisiae)
- Y-tubulin small complex bound to two alpha/beta tubulin dimers (Cellular component from Saccharomyces cerevisiae)
- Unknown protein (5 kDa, Protein from Saccharomyces cerevisiae)
in situ subtomogram average of C. elegans microtubules in mitotic centrosomes
Tollervey F, Rios MU, Zagoriy E, Woodruff JB, Mahamid J
bioRxiv (2024) [ DOI: doi:10.1101/2024.04.03.587742 Pubmed: 38617234 ]
in-situ subtomogram average of C. elegans centrioles in centrosomes
Tollervey F, Rios MU, Zagoriy E, Woodruff JB, Mahamid J
bioRxiv (2024) [ DOI: doi:10.1101/2024.04.03.587742 Pubmed: 38617234 ]